
Then for PKASY=1, you will get CEps*dCObsdPKASY1 as the second cv after fitting. This is the assumption they share the same cv.
Observe(CObs = (C * (1 + CEps*dCObsdPKASY1*(PKASY=1) ))) This should change to me only the value of CEps. What you want to say is that the cv is different for the 2 assays. Stparm(V = tvV * (WTBL/70)^dVdWTBL * exp(dVdDIAL1*(DIAL=1)) * exp(nV))
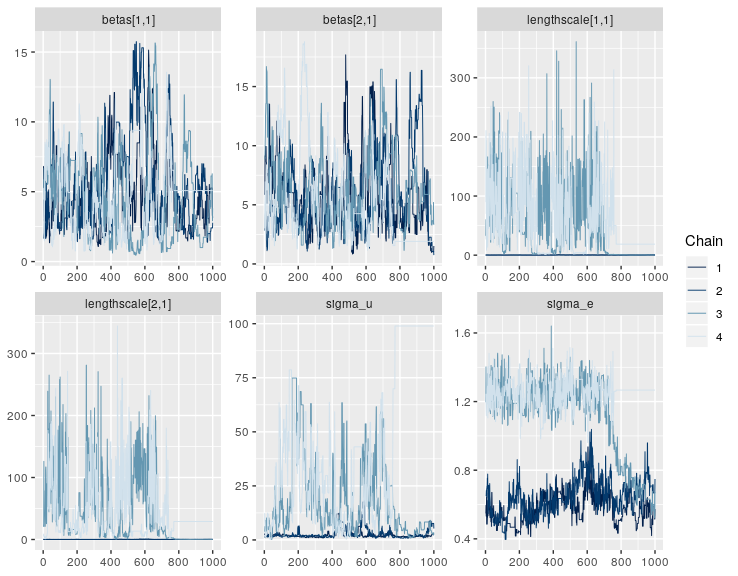
#Nonmem 3 new residual variability models eta on sigma code
We are getting error of around 40% in NONMEM, so I know I am making some error here.Ĭan you please check below mentioned code and suggest if I am making any error in observe statemetn? I am able to run the model with below mentioned code however, I am getting really high estimates for proportional error variance ~200. I want to add assay type (categorical with values 0 or 1) as a covariate on proportional residual errors.
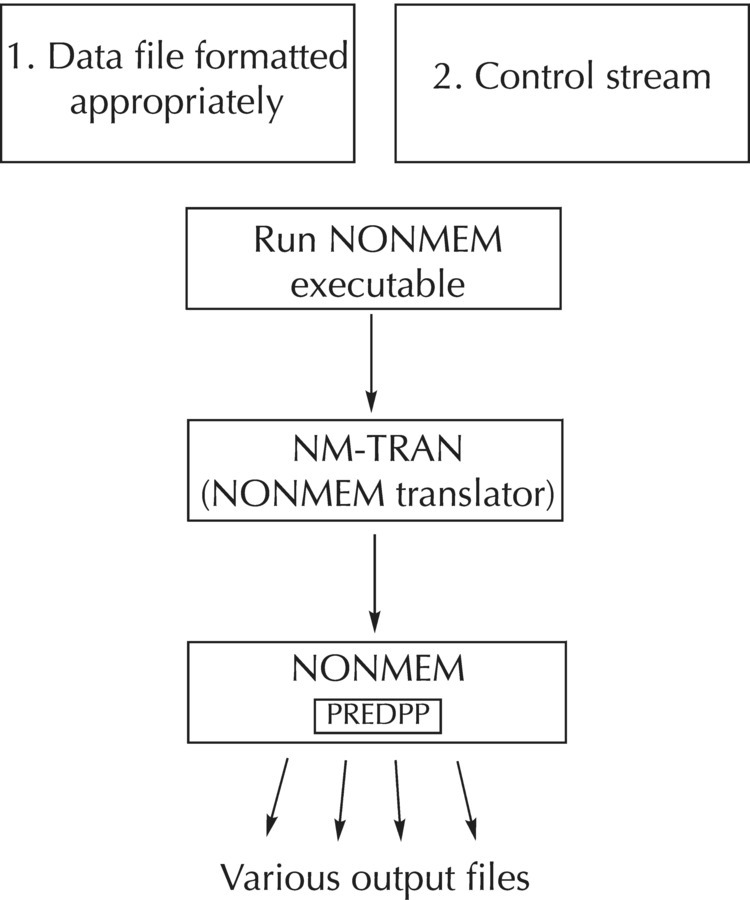
I am fairly new to PML coding and am trying to reproduce a pop PK model in Phoenix NLME that we had earlier run in NONMEM.
